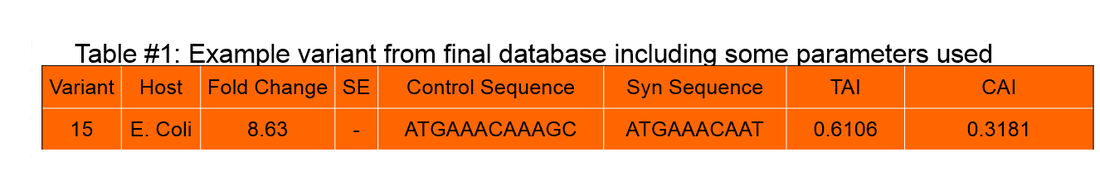
Database

Nupack software was used to calculate the Gibbs free energy of the reaction, an input which was needed in iSCOOP
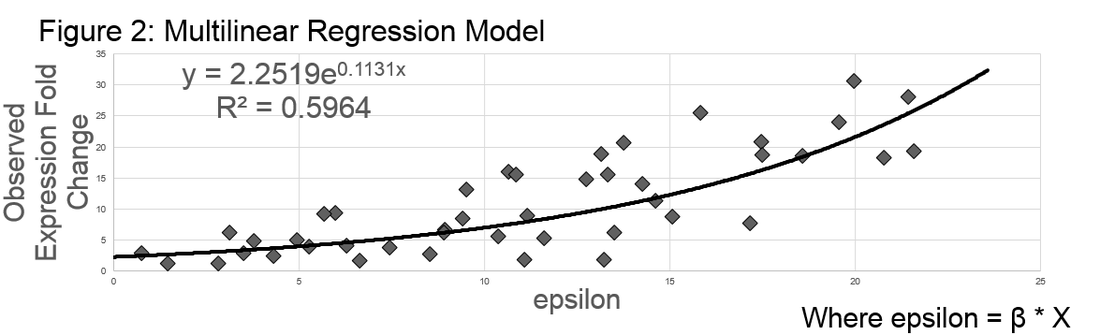
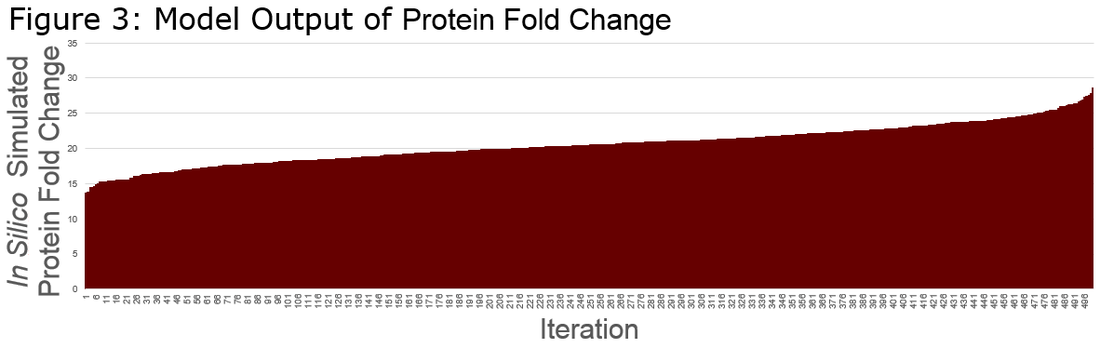
Model
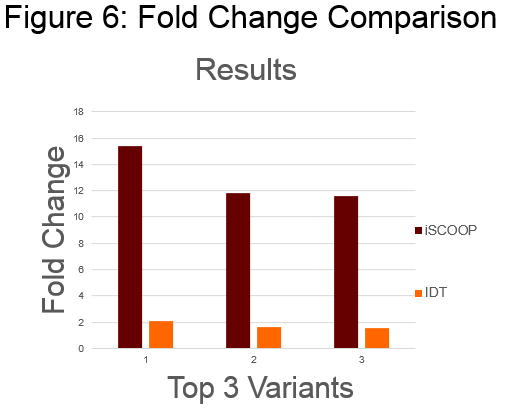
Validation
Fig. 4: The model's in silico predictions correlate with an R-squared 0.6646 to our observed increases in protein expression. The model overestimates the fold-increase, as the ratio of observed to predicted fold-increase is 0.757.
 The five most influential parameters can be seen in this table.
The five most influential parameters can be seen in this table.